It can quantify antibody fluorescence intensity, colocalization, cell density, or even track cells through time-lapse movies. Cell Profiler/Cell Analyst – Image Batch Processing and Data AnalysisĬell Profiler can extract quantitative measurements from thousands of images through a custom pipeline that can first process and then analyze your images. The select grouping of plugins is geared for neuroscience.Ģ. There were a few plugins that were only available in FIJI. Downloading FIJI instead of ImageJ just starts you out with more base plugins so you don’t have to hunt around and add them to your ImageJ program. It can also easily handle 3D stacks of confocal microscopy images, and perform complex quantitative analysis.įor more information visit their website: įIJI (FIJI Is Just ImageJ) is a bundle of the top plugins available for ImageJ. It can do simple things like crop, label, and alter the brightness and contrast of fluorescence images. Being proficient at using ImageJ is essential for most image processing and analysis. Download it, search through the plugins to see what’s available and test them out. ImageJ should be the first program you become familiar with when looking for image analysis software. 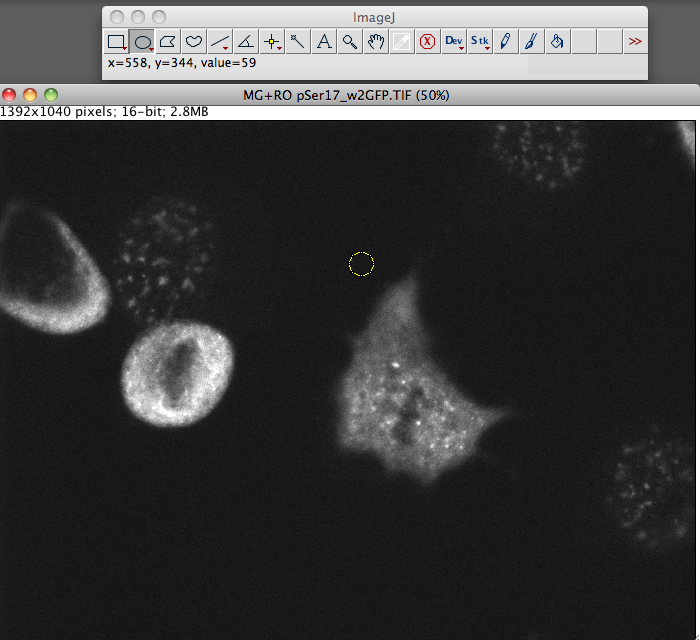 
Here are my Top 5 favorite time/life savers: It took me years (actually!) to find some of them, so I thought I would pay-it-forward by sharing them with you! Kudos to everyone who takes the time to create these free open source programs. Out of sheer necessity to analyze a massive image set before I turned grey, I scrounged around for free open source software programs to help analyze my confocal microscopy image stacks. I hope this was somewhat helpful, and maybe others can pitch in their thoughts as well.If your lab was like mine was, you struggle to afford new antibodies, let alone expensive image analysis software programs like Neurolucida to analyze your data. I don’t really think there’s a way around that though, except by just making sure all of the step sizes are equivalent for the various z-stacks you compare (so if it’s happening in one, it’s happening in all of them). This is because he felt like it was possible for the same molecules to show up as fluorescence in more that one slice, and thus their colocalization could count double or triple. Someone from Nikon told me though that they felt it was important to have the incremental steps in the z-stack be the same for all of the z-stacks being compared (so even though one might be 30 slices and another 45, the step size for both should be the same). So a single slice is just like one quadrant of an image, but the value given is for the image in its entirety, if that makes sense. What I determined from all the sources I read and people I talked to was that the JaCOP plugin is NOT calculating an average, but instead just giving the coefficients as proportions relative to the total amounts of fluorescence that is there within the entire z-stack as a whole. Sorry it took me a couple days to see this and respond. I still can’t find the exact answer I’m looking for though… I have read that paper as well as a few others over the past few months, though I have to admit some of the information goes over my head. I have to give my thesis seminar pretty soon, and I’m afraid someone on my committee will end up asking me just how exactly these values are calculated from a z-stack…specifically if they are an average or not, but I just can’t seem to find that information anywhere! Thank you for the paper suggestion. The ones I have now range from about 35 to 50 slices. I’m glad to hear you think it is acceptable!īut since all of my images are z-stacks, I was trying to figure out how important it is for them to all be the exact number of slices. I’m trying to be as objective and consistent as possible in setting the threshold values, but I was also unsure just how much that is frowned upon. It seems like most of the papers I’ve read strongly caution against setting your own threshold values though, and instead advocate using the Costes method…but every time I’ve used that method it sets the threshold values way too low. The process you are describing sounds like what I have been doing with the JaCOP plugin which allows me to set my own threshold values and get a Manders M1 and M2 coefficients (red coloc/ total red and green coloc/ total green, which I think is the same as what you described, right?). Thank you for your reply! I agree about the intensity-based colocalization methods, which is also why I chose the Manders method because it was my understanding that it ignores intensity differences…but someone correct me if I’m wrong. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
